In mid-January, a group of computer scientists and biologists from the University of Vermont, Tufts, and Harvard announced that they had created an entirely new life form — xenobots, the world’s first living robots. They had harvested skin and cardiac cells from frog embryos, designed and sculpted them to perform particular tasks with the help of an evolutionary algorithm, and then set them free to play. The result— it’s alive! — was a programmable organism. Named after the African clawed frog, Xenopus laevis, from which they were harvested, these teams of cells were wholly liberated from constraints of frog DNA. Their behavior was determined by their shape, their design. They can already perform simple tasks under the microscope. And due to their minuscule size and great adaptability, they could soon be put to work in the human body to deliver new medical treatments, or out in the world to do environmental clean-ups.
Naturally, this raises all kinds of questions and challenges for both humanity at large and synthetic biology. Earlier this year, Grow’s Christina Agapakis discussed some of them with Michael Levin, one of the scientific minds behind xenobots. This mind-bending conversation delves into how Levin and his team discovered and conceptualized a new life form, the possible applications they envision, and what this all says about the different ways computer scientists and synthetic biologists think about DNA.
Christina Agapakis: I’m interested in the language you use to explain these xenobots — how you conceptualize what some people are calling a new life form. What inspired this project? What kicked off this research direction?
Michael Levin: Our group studies cellular decision-making. We’re interested in what is called basal cognition: the ability of all kinds of biology — from molecular networks to cells and tissues, from whole organisms to swarms — to make decisions, learn from their environments, store memories. We were thinking about new forms of minimal model systems that are synthetic or bio-engineered, where we could build basic proto-cognitive systems from scratch and really have the ability to understand where their capacities come from.
We are also interested in the plasticity of cells and tissues. What can they do that is different from their genomic defaults? In our group, we think that what the genome does is nail down the hardware that these cells have: the proteins, the signaling components, and the computational components. When these things are actually run in multi-scale, biological systems, there’s an interesting kind of software that drives the structure and function. We’ve been working for years now on the reprogrammability of that software — the idea that you can give cells and tissues novel stimuli or experiences and thus change how they make decisions in terms of morphogenesis, in terms of behavior, without actually changing their hardware.
Christina: That’s fascinating because it’s different from how the synthetic biology world typically uses words like hardware and software. When folks at Ginkgo, or other molecular or synthetic biologists, talk about software, they often mean the DNA code. What do you mean when you say software?
Michael: I have a completely different perspective. I don’t think DNA is the software. Not that only one metaphor is valid, but the one that we have found useful is this idea that the real-time physiology of the organism is the software. My background is in computer science, so I come at this from more of a computational perspective. The important thing about software is that if your hardware is good enough — and I’m going to argue that probably all life at this point is good enough — then the software is rewritable. That means you can greatly alter what it does without meddling with the hardware.
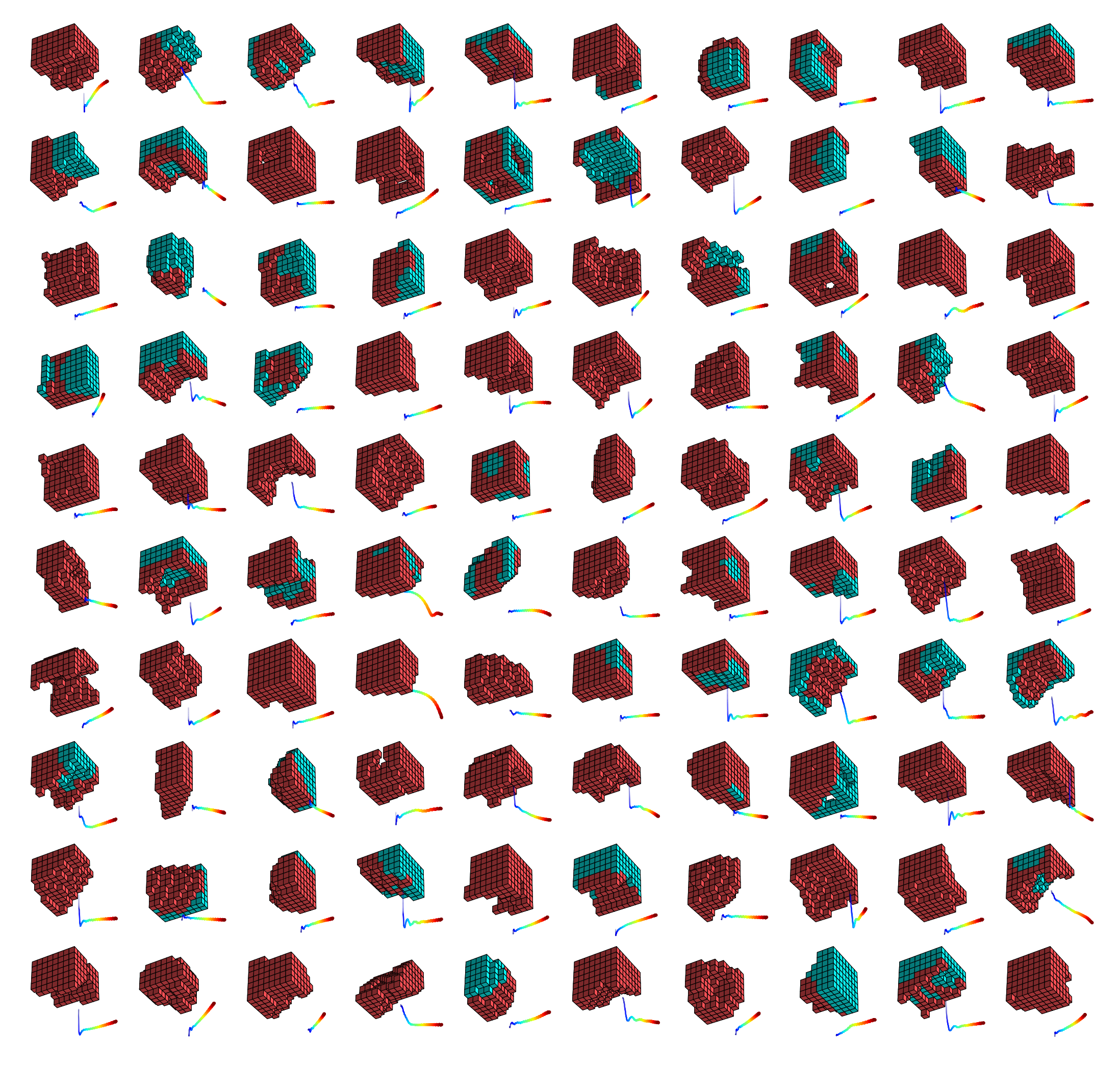

People are very comfortable with this in the computer world. When you switch from Photoshop to Microsoft Word, you don’t get out your soldering iron and start rewiring your computer, right? In fact, that is exactly how computation was done in the 1940s, and I think that’s where biology is today. It’s all about the hardware. Everybody’s really interested in genomic editing and rewiring gene regulatory networks. These are all important things, but they are still very close to the machine level. In our group, we think of the DNA as producing cellular hardware that is actually implementing physiological software, which is rewritable. That means you can greatly alter the behavior of the system without actually having to go in and exchange any of the parts. I think that’s the amazing thing about biology. The plasticity is quite incredible.
I’ll give you a simple example of what I mean by software. We have these flatworms, planarians, and they have a head and a tail. What we’ve shown is that there’s an electric circuit that stores the information of how many heads the planarian is supposed to have and where the heads are supposed to be. The important thing about this circuit is that it has this interesting memory aspect. We can transiently rewrite the stable state of this electric circuit with a brief application of ion channel drugs, inducing a permanent change in the animal’s body plan.
Christina: It’s actually electric.
Michael: It’s physically electric. A large chunk of our lab works on developmental bio-electricity — electrical communication and computation in non-neural tissue. We’ve discovered that the planarian’s tissues store this steady state electrical pattern, which dictates how many heads they are supposed to have. Their genetics give you a piece of hardware, which self-organizes an electrical pattern that causes cells to build one head, one tail. That’s the default. Now that we understand how this works, and it took a good 15 years to get to this point, we can go in and rewrite that electrical state. The cool thing about that circuit is that, like any good memory circuit, once you’ve rewritten it, it’s stable, it saves the information, unless you change it back.
You can rewrite it so that, if you cut the worm in half, it creates worms with two heads, no tail. Those two-headed worms continue to make two-headed worms, if you cut them in half again. All of these worms have completely normal genomic sequences. We haven’t touched the DNA at all. The information on how many heads you’re supposed to have is not directly nailed down by the genetics.
People are comfortable with this in electronics: you turn it on and it does something. If it’s an interesting piece of electronics, you can reprogram it to do something else. It turns out that biological electric circuits are exactly like this. They have incredible plasticity. They are very good at enabling some of these electrical states to be rewritten. Once we know how to do it, you can actually reprogram those pattern memories. These are structures in the tissue that tell the cells what to do at a large scale. That’s the kind of thing I mean by software. There are physiological structures in the tissue that serve as instructive information for how it grows.
Christina: And how did this research lead to xenobots?
Michael: We wanted a synthetic, bottom-up system where we could see this from scratch. In particular, we wanted to understand not just the hardware of cells, but the algorithms that enable them to cooperate in groups. Josh Bongard at UVM is a leader in artificial intelligence and robotics. He and I have had many conversations about how biology can inform the design of adaptive and swarm robotics and AI, and how the tools and deep concepts from machine learning and evolutionary computation can help understand different levels in biology. He was a very natural partner for this work, and we decided to establish a tight integration between computational modeling and biological implementation.
We took some cells from a frog embryo, and we let them re-envision their multicellularity. Individual cells are very competent. They do all sorts of things on their own. How do you convince them to work together toward much bigger goals? Cells in the body work on massive outcomes, things like building a limb, or building an eye, or face remodeling. They are called upon to do it and they stop when it’s done. So, when a salamander loses a limb and the limb is rebuilt, you can see that this collective knows what a correct salamander looks like, because it stops all that activity as soon as the limb is complete. We wanted to know: how do cell collectives store and process these kinds of large-scale anatomical pattern memories?
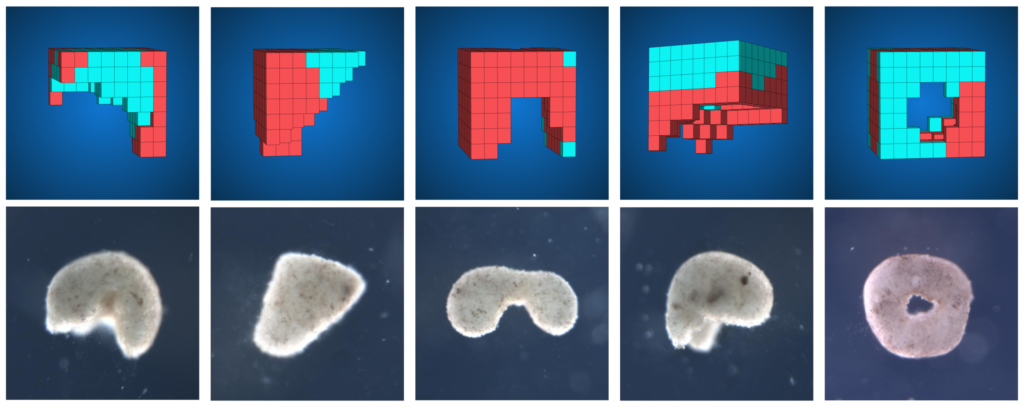
We liberated the frog cells from the boundary conditions of the embryo. And we said to these cells: here you are, in a novel circumstance, in a novel environment: what do you want to do? What we observed is that the cells are very happy to once again cooperate with each other. They build a synthetic living machine with behaviors that are completely different from what the default would be. They look nothing like a tadpole, nothing like a frog embryo. We have lots of videos of them doing interesting things. They wander around, they do mazes, they cooperate in groups, they communicate damage signals to each other. They work together to build things out of other loose cells.
I think what we’re scratching at here is just the beginning of understanding what cells are willing to do under novel environments, and how plastic they really are. That was the origin of this project: to try to understand what cells can do beyond their default group behaviors. That’s what we have here.
Christina: You’re collaborating with philosopher/ethicist Jeantine Lunshof on the next phase of this project. What are the ethics of manipulating life in this way?
Michael: In terms of the ethics of the platform itself, I don’t find it particularly far past the range of things that have already been done. We [as a society] manipulate living cells all the time. We have a food industry where we manipulate whole organisms, adult mammals, who we can be quite certain have some degree of agency. In that sense, I don’t think these things push the envelope on any ethical issues. I do think that they highlight the inadequacies of the definitions that we throw around on a daily basis. There are all sorts of terms we think we understand, people use them all the time: animal, living being, synthetic, creation, machine, robot. Lots of work in robotics and synthetic biology has been showing us recently that we actually don’t understand what those things are at all.
I gave a talk once and I referred to a caterpillar as “a soft-bodied robot” and some people complained about that. They said, oh my God, how can you call it a robot? That’s because they’re thinking of robots from the ‘60s — you know, these things that are on the car assembly line. That’s a very narrow and, in 2020, not very helpful view of what a robot actually is. We are still trying to come up with good definitions for all these things. What do we really mean when we say machine? If you think some things are not machines, what does that mean? And what do you think they have that makes them different from machines? This raises very interesting philosophical questions.
Christina: That’s part of the work your xenobots are doing — philosophical, conceptual, perhaps even artistic work. I’d like to talk a little more about the practical uses and implications. What types of applications do you imagine? If we’re in the 1940s or 1950s of biology: how do these look 50 years from now?
Michael: I would start with what I would call near-term applications. You could imagine these things roaming the lymph nodes and collecting cancer cells, sculpting the insides of arthritic knee joints. You could imagine them collecting toxins in waterways. Once we learn to program their behavior, which is the next thing that we’re doing, you could imagine a million useful applications, both inside the body and out in the environment.
This is the kind of model system that can tell us a lot about where true plasticity comes from. What are the strategies that you might want to build into your robots or your algorithms that would allow them to respond to novelty the way that living cells in collectives do?
Individual cells are very competent. They do all sorts of things on their own. How do you convince them to work together toward much bigger goals?
MICHAEL LEVIN
The medium-term applications, I think, are more in the fields of Regenerative Medicine and AI, specifically looking at how to program collectives. If you are going to rebuild somebody’s arm, or a limb, or an eye, or something complex like that, I think that we are going to have to understand how cells are motivated to work together. Trying to micromanage the creation of an arm from stem-cell derivatives, I think, is not going to happen in any of our lifetimes.
We need to understand how you program swarms to have the kinds of goal states that we want them to have, so they can build organs in the body, or for transplantation, or for complex, synthetic living machines. If we figure out how this works, and where these anatomical goal states come from, we will be able to make drastic improvements in regenerative medicine. It’s not just that we might have these bots running around our bodies. It’s the fact that we will be able to program our own cell collectives at the anatomical level, not at the genetic level.
The idea is to understand how cell collectives encode what it is that they’re building, and how you could go about rewriting this — to offload the computational complexity onto the cells themselves and not try to micromanage it. Our recent paper on frog leg regeneration is a good example of that. Frogs normally do not regenerate their legs, unlike salamanders, and we figured out how to make them do it. The intervention is 24 hours, and then the actual growth takes 13 months. So it’s a very early signal. We don’t hang around and try to babysit the process. You figure out how to convince the cells what they should be doing and then you let the system figure out how to do it on its own.
Christina: Maybe it’s an accident in the history of biology, but I think our narratives are so dominated by the idea of molecular control — by DNA as this central driver of everything. There are synthetic biologists who basically say that DNA is the only thing that matters. You’re highlighting a very different perspective. You’re saying the stimuli plus the cell and its existing hardware can create a totally different outcome. That has fascinating implications for how we think about genetic determinism in synthetic biology, and, more generally, in human behavior, in human outcomes, and so many other different things.
Michael: I think we have a good framework to think about this, which is the progress in computer science. I am not saying that living things are computers, at least not like the computers you and I use today. What I mean is: computers are fundamentally a very wide class of devices that are in an important sense reprogrammable. My point about DNA is that when people think about genetic determinism, they’re not giving the DNA enough respect. DNA doesn’t give you hardware that always does the same thing and is determined. DNA is amazing. DNA has been shaped by evolution to produce hardware that is eminently reprogrammable. I think we need to respect the fact that evolution has given us this amazing multi-scale goal-driven system where the goals are rewritable.

I’ll give you another very simple example. Tadpoles need to become frogs. And tadpole faces need to be deformed to become frog faces: the eyes have to move, the jaws have to move, all this stuff has to move around. It used to be thought that what the genetics encodes during metamorphosis is a set of movements that would make that happen. If every tadpole looks the same, and every frog looks the same, then that works, right? You move the eyes, you move the mouth, everything moves a prescribed distance in a certain direction, and you’re good. To test this, we made what we called Picasso tadpoles. Everything is in the wrong place: the mouth is up here, the eyes are sideways, the jaws are displaced, everything is just completely moved around. What we found is that those tadpoles still become pretty normal frogs. Everything moves around in really unnatural paths, and they keep moving until a normal frog face is established.
This shows that the genetics don’t just give you a system that somehow moves everything in the same way every single time. They give you a system that encodes the rough outline of a correct frog face, and then it has this error minimization scheme where what the hardware does is say, wherever we’re at now, I’m going to keep taking steps forward to reduce that error to as low a level as possible. That’s just an example of this goal-directed plasticity. And I think it’s important to realize that that’s the beauty of the hardware that evolution has left us with. That’s the trick. The hardware is much more capable than we’ve been giving it credit for.
Christina: Hearing you talk about these non-genetic factors, it does seem like you’re pointing at what’s fundamentally missing from how we think about synthetic biology and genetics more broadly. There’s the critique of genetic studies of human behavior that basically says: you can never really control for all the variables — social, environmental, whatever — so your studies of the genes for almost anything are going to be fatally flawed. Then conversely, if you are only seeing a small picture of what it means for a human to be anything — healthy, smart, athletic, beautiful — engineering someone’s DNA with those outcomes in mind is probably not going to get you what you want, and we should focus on the ways we can change a person’s environment that will make them healthier, smarter, happier etc. If biology is so plastic, why do you think the genetic idea and the dream of designer babies persists?
Michael: I want to be clear: I’m not denigrating genetics in the slightest. I think understanding the hardware is critical. You’re not going to get very far without understanding your hardware. And I think that genetics is essential for that. In some cases, working at the level of hardware is fine. If you want to fix a flashlight, you can do everything you need to do at the hardware level. Or if people want to make, say, bio-engineered bladders, a sphere, you might get away with literally micromanaging this thing directly with some stem cells and growing them on a scaffold. But if you really want to make large-scale control of complex anatomy, I would say there’s not a compelling history of capabilities today that would suggest that we have good anatomical control at the genetic level. We are not very good at it right now.
A lot of people make a promissory argument. They say, turn the crank, keep going, we’re going to sequence a whole bunch more stuff. We’re going to do a lot more transcriptomics. We’re going to do a lot of genomics. We’re going to keep at it and someday we’ll just be able to do it. You want seven fingers? You got it. You want gills? Fine. I don’t find that promise very compelling.
You could spend all your time drilling down into the molecules that it’s made of, but at some point, you have to ask yourself, what are they in a cybernetic sense? What’s the function of this thing? What are the control loops? What are the internal capabilities? Is it reprogrammable? Is its structure modular? All of these things are completely invisible at the level of the hardware. I think it would be incredibly unwise to throw away all the lessons of engineering, of cybernetics, of computer science. Can you imagine where our information technology would be today, if every change you wanted to make you had to make at the hardware level, or even in machine code? I mean, it’d be insane. We wouldn’t have anything.
DNA has been shaped by evolution to produce hardware that is eminently reprogrammable.
MICHAEL LEVIN
Christina: I do molecular stuff. I live in that world. I’m interested in your critique or challenge of that. Maybe there is a limitation to how we’re imagining things. You might be able to sort of open up much more interesting questions and possibilities if you’re looking at how things grow. How did you arrive at some of these more holistic perspectives?
Michael: Some of this stuff is absolutely ancient. I mean, back in the ‘40s, you have this biologist who took a paramecium, or a similar kind of single-cell animal, which is covered with these little hairs that all point the same way. He took a little glass needle, cut a little square into the surface of the thing, turned it 180 degrees, and put it back. An amazing technical feat. Now the hairs are pointing the wrong way in that little square. And what he found is that when the paramecium divides and has offspring, all of the offspring now have little squares of hair that are pointing the wrong way. Why is that? It’s because the structure of the cortex is templated onto the previous one. So when it makes a daughter cell, it just copies whatever it has. The non-genetic piece of information is critical. This is the original demonstration of true epigenetics. This information is simply not in the genome.
In our view, turning on and off specific genes to make specific cell types is such a tiny corner of all this. It’s critical, but we’ve got to think more broadly, in pattern control, in large scale goal-directedness, and think about the computations that all these different levels are doing to get where they’re going. There’s no way we’re going to do what we need to do in biology without an appreciation of those other levels.
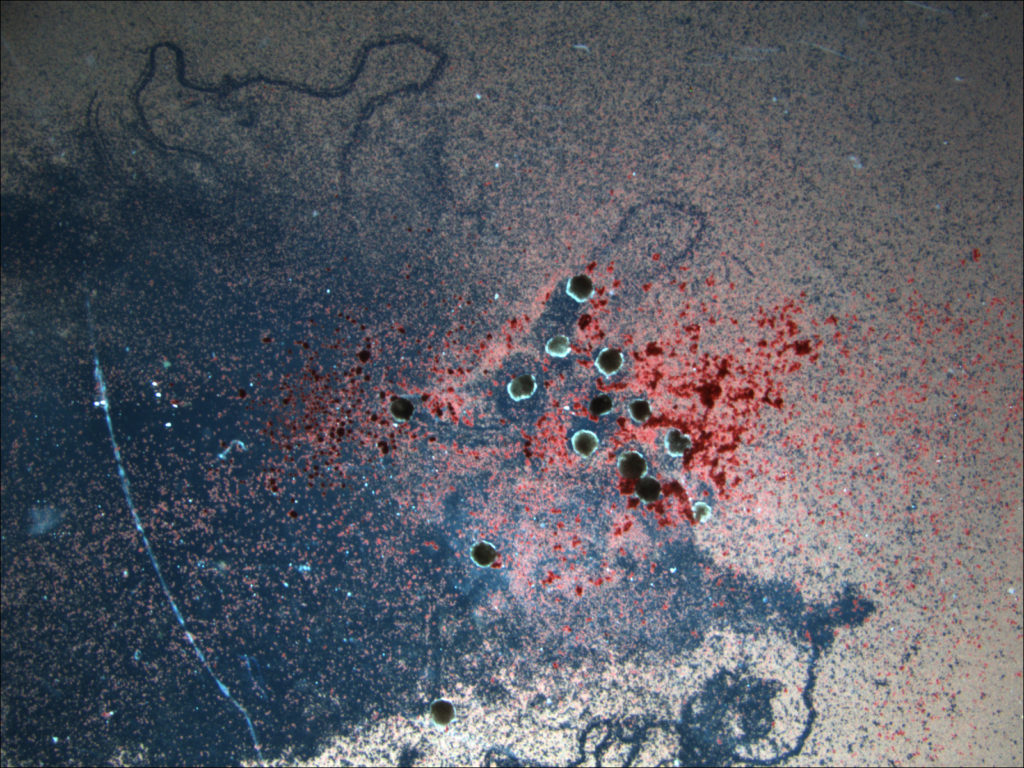
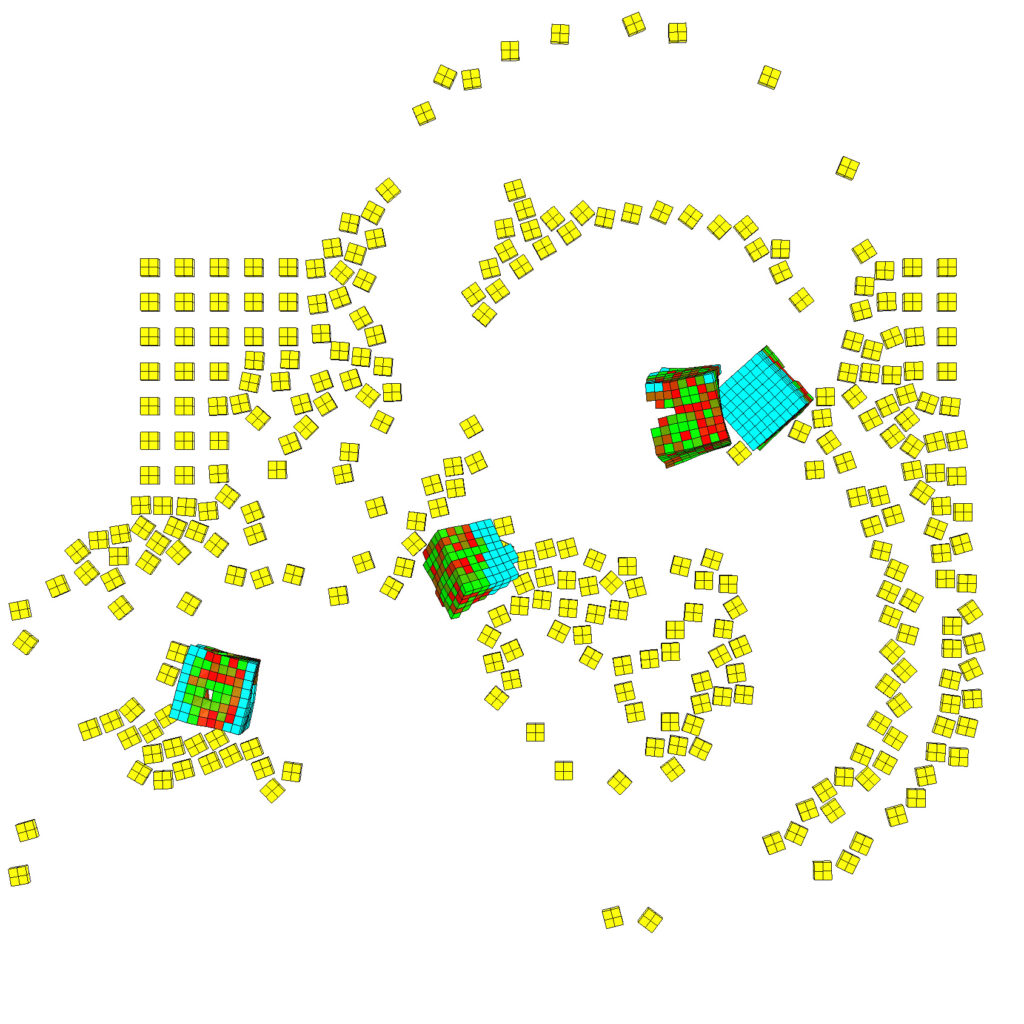


(Left) Five red-cyan designs are placed amid a lattice of simulated debris, in yellow (Right) The traces carved by a swarm of these organisms as they move through a field of particulate matter. (Credit: left, Sam Kriegman; right, Douglas J. Blackiston)
Christina: One thing we’re interested in doing in Grow is outlining future scenarios, the possibilities for biology. What’s the end goal for xenobots?
Michael: I’ll jump way forward because it’s more fun that way. Regeneration and limb regeneration is critical, but we can fantasize further than that. In the sort of asymptote of all of this, I see two things that I think should be possible. On the one hand, I think this is progressing towards a total control of growth and form. At some point, when we really know what we’re doing — when we actually know how morphology is handled — you will be able to sit down in front of a kind of a Computer Aided Design (CAD) system, and you will be able to draw whatever living creature you want, in whatever functional anatomy you want. It might be for something with an application here on Earth. It might be an organ for transplantation. It might be a creature that you’re going to use in colonizing some far off world. Whatever it’s going to be, you are going to be able to sit down and specify at the level of anatomy, the structure and function of a living creature at the high level, and then this will sort of compile down and let you build the thing in real life.
Right now, we can only do this in a very simple set of few circumstances, but ultimately if we really knew how this worked, we would be able to have complete control. People talk about constraints on morphogenesis and on development. I think those are constraints on our thinking, not on the actual cells themselves. I think you should be able to build pretty much anything within the laws of physics. Almost anything.
Christina: You talk about not wanting to micromanage the cells, but you’re sort of shaping the growth, you’re facilitating and influencing it. What do you mean when you say control?
Michael: To use another super-anthropomorphic phrase, I think our goal is to convince the cells to do what we want them to do. Your goal is to exploit the computational capacity of the system, and to understand how it is that you communicate your goals to the growing tissue. We’re already starting experiments on basically behavior-shaping the tissue with rewards and punishments. Humans have figured out over 10,000 years ago that we don’t have any idea how the animal works inside. But what we do know is that if you give it rewards and punishments you can achieve outcomes that you like. This ability of living things to change their behavior, to make their world better, is ancient, and that’s how rewards and punishments work. This probably works all the way down. Your goal is to convince the system to do what you want it to do not to try to build it up brick by brick.
It’s a top down view of control. Your goal is to specify the end goal, and let the system figure out how to get there. That works very well with systems that have the necessary IQ. So that might work with your kids, and it might work with various other animals. It doesn’t work real well with a cuckoo clock. It just doesn’t, because its hardware system isn’t amenable to that kind of control. As always in science, you have to figure out when these kinds of approaches are appropriate and when are they not. There’s a large class of systems where that’s useless, and a massive class of biological systems where I think that’s going to be the way to go. I think that’s part of our future.
One thing I think this is showing us is that focusing on the brain as the source of inspiration for machine learning is derived from a very specialized architecture. I’ve been suggesting that a true general purpose intelligence is much more likely to arise not from mimicking the structure of the core of the human cortex, or anything like that, but from actually taking seriously the computational principles that life has been applying since the very beginning.
Christina: Paramecia?
Michael: Even before that. Bacteria biofilms. All that stuff has been solving problems in ways that we have yet to figure out. They’re able to generalize, they’re able to learn from experience with a small number of examples. They make self-models. It’s amazing what they can do. That should be the inspiration. I think the future of machine learning and AI technologies will not be based on brains, but on this much more ancient, general ability of life to solve problems in novel domains.
Christina: Bacterial intelligence.
Michael: Exactly. And not just bacterial — individual cells building an organ and being able to figure out how to get to the correct final outcome from different starting positions, despite the fact that you went in there and mixed everything around. I think a lot of our true general AI in the future is going to come from this sort of work on basal cognition. I guess my theme is consistent. I think we need to step back from any kind of uniqueness of the human condition, and try to generalize it more and more broadly. We need to take evolution seriously.
This conversation is Part 1 of Grow’s coverage of xenobots. Continue reading Part 2, “The Lab Philosopher“, where we interview Harvard ethicist and philosopher Jeantine Lunshof on the long-term ethical and the philosophical implications of the discovery.
CONTACT THE RESEARCHERS:
Michael Levin
Director, Allen Discovery Center at Tufts University
Associate Faculty, Wyss Institute at Harvard University
drmichaellevin.org | @drmichaellevin