I. In the Shallows
Looking out at sea, suffering yet another bout of motion sickness, Charles Darwin saw nothing but a barren “desert of water.” How wrong he was would only become clear decades later with the birth of modern oceanography. Forty years after Darwin’s voyage on the HMS Beagle, another British naval ship, the HMS Challenger, was retired from colonial military service and sent out on a scientific expedition to figure out what the hell was going on under the sea. Cannons and ammunition were removed to make space for laboratory instruments, jars awaiting specimens, and hundreds of miles of rope. And so, in 1872, a crew of English and Scottish naturalists set sail on a voyage around the world, sampling the bottom of the ocean and pulling netfuls of alien creatures into daylight. Far from a water desert, they found the richest ecosystems on Earth.
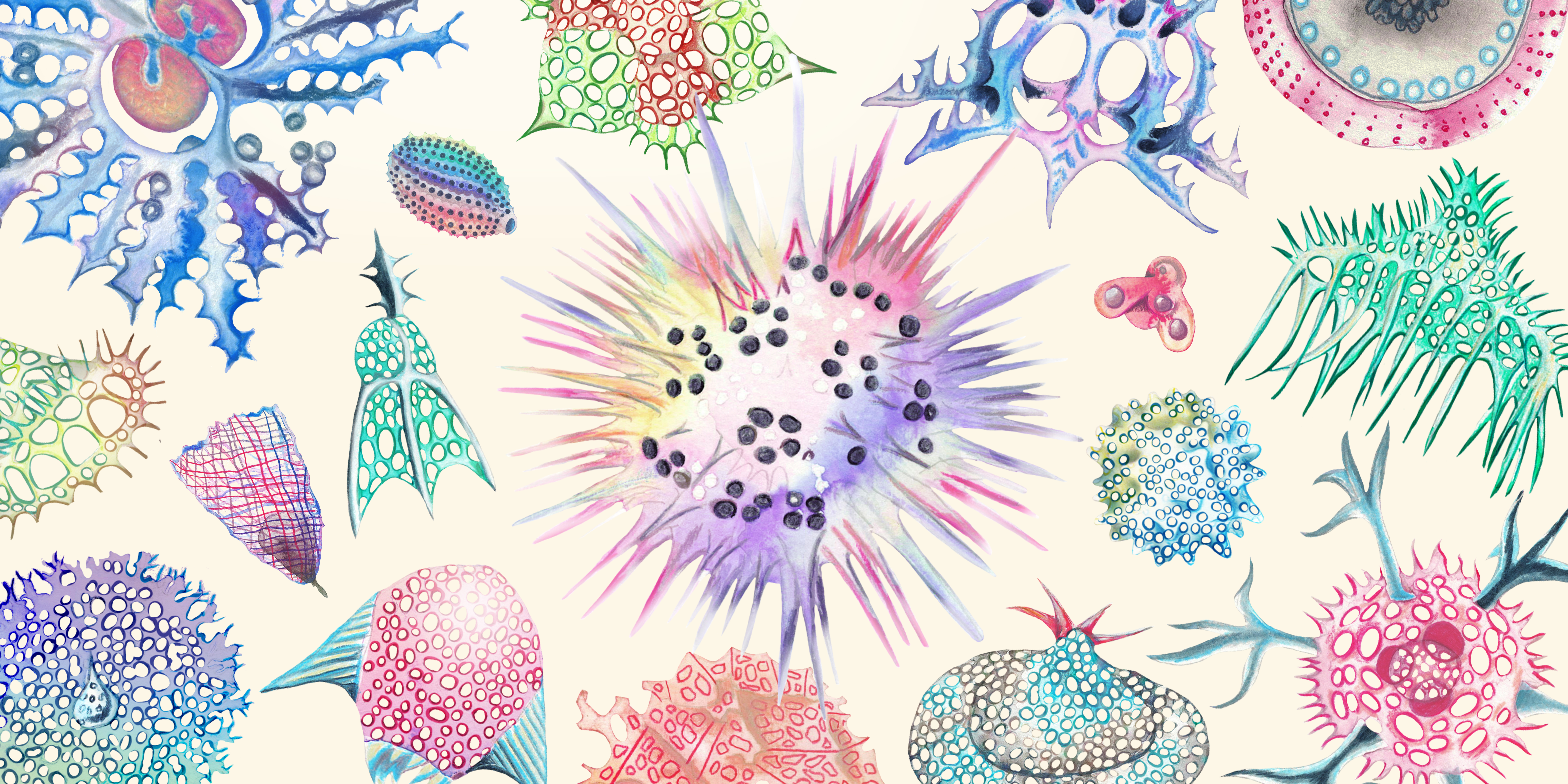
Among all this vastness, some of their most remarkable discoveries would be very small. Microscopic, in fact: like the tiny glass skeletons that had been collecting on the ocean floor for millennia, remnants of plankton known as Radiolaria. To sift through these spoils, the Challenger crew spread them among a pan-European roster of scientists. That’s how zoology professor and illustrator Ernst Haeckel got his hands on four thousand new species of radiolarians, which he spent a decade documenting, classifying, and sketching. His drawings — some of the most startlingly beautiful scientific illustrations ever made — captured these fabulously patterned and twisted houses of glass. In the nineteenth century, it was hard to believe that single cells could function as entire organisms, let alone create delicate and microscopic globes, discs, thimbles, thistles, and spiked bodies.
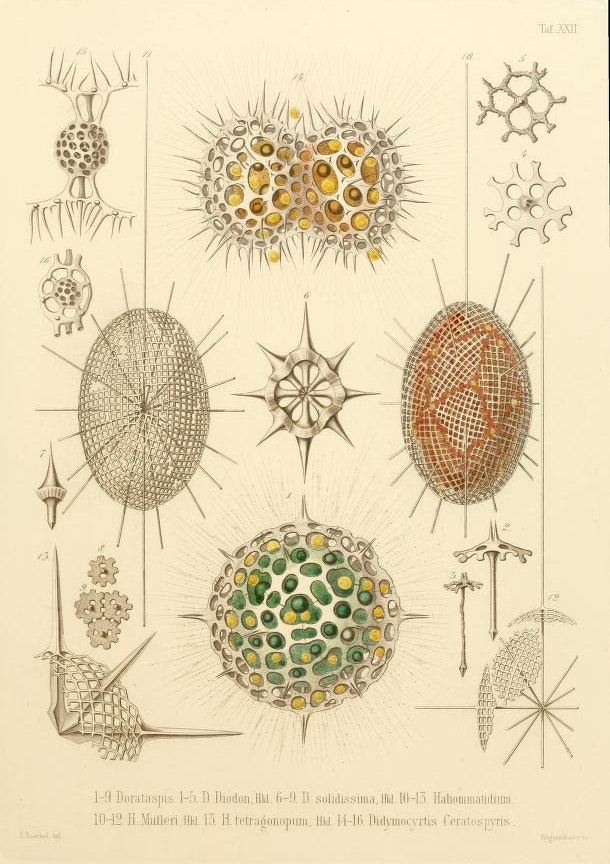
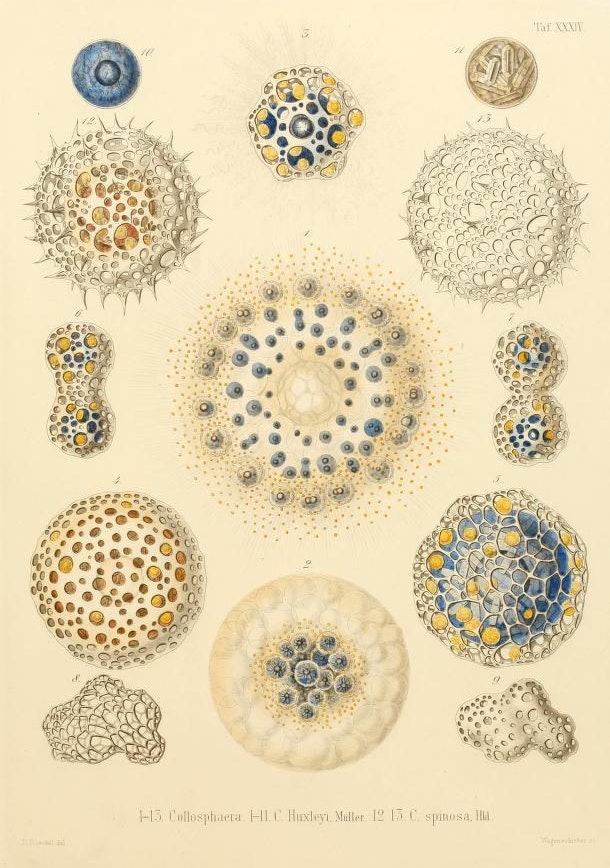
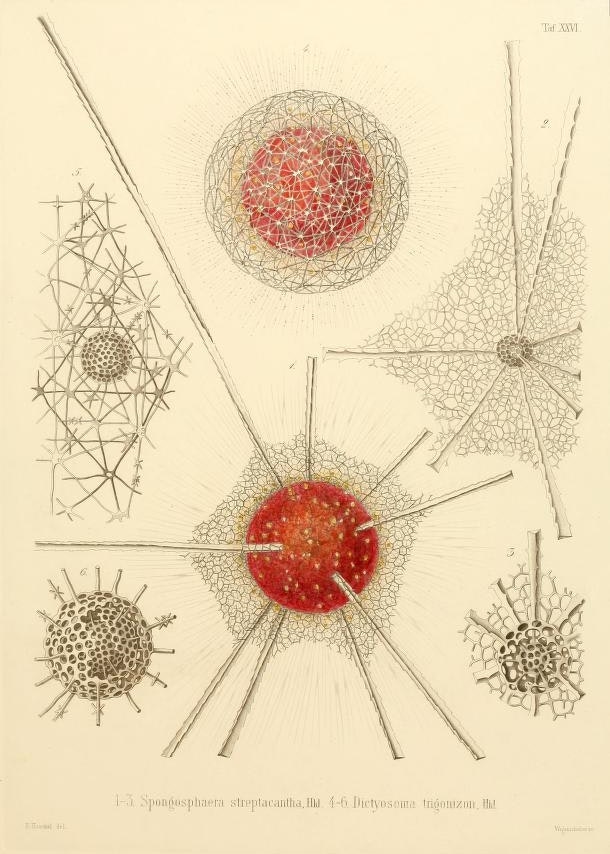
Haeckel proposed the category Protista, meaning “the first of all,” or “primordial,” to describe unicellular organisms like Radiolaria that were neither plants nor animals, despite sometimes sharing their traits. Today, seaweed, slime molds, and the malaria parasite Plasmodium are all considered protozoa. If you aren’t familiar with these protists, there’s a reason why. More people today study sparrows than all marine protists. A poorly understood and risky area of research, they remain the “dark matter” in the tree of life. What potential is slumbering in this great unknown?
Imagine if we achieve today’s vision for synthetic biology. Genetic circuits become the backbone of modern medicine, every commodity is fermented into existence, alternative protein products abound in every aisle. Even then, protists will remain a treasure trove of inspiration. They are a neglected component of the microbial life that surrounds and saturates us. We have barely even scraped the surface of their genomes. Until we can comprehend the invisible majority of eukaryotic life, we are playing in the shallows of an ocean of biological potential.
II. The Tree of Life
In 1969, Robert Whittaker proposed five kingdoms to classify all life. Prokaryotes formed the root of this tree, with three branches emerging for plants, animals, and fungi and protists at the forks. The last kingdom, Protista, was a catch-all category for organisms like the amoeba: taxonomic leftovers that seemed to belong more with each other than with any other group. This classification followed from our experience of the natural world: we encounter animals, plants, and fungi every day. Organisms that elude our senses default as something other. Luckily, we’ve learned a lot in the past 50 years since Whittaker, including how to sequence organisms’ DNA and deduce their evolutionary relationships.
Until we can comprehend the invisible majority of eukaryotic life, we are playing in the shallows of an ocean of biological potential.
The latest revision of the tree of eukaryotes prioritizes clades, groups that contain all the organisms descended from one common ancestor. Whittaker’s three branches have been split into twenty-eight clades. None of them are entirely plants, animals, or fungi, and they all have a piece of the defunct Kingdom Protista. Land plants and green algae share one branch. A second is home to animals, fungi, and several related kinds of protozoa. That leaves twenty-six lineages representing mostly unicellular, aquatic, soil-dwelling, or parasitic organisms that are more distantly related to each other than humans are to yeast (a kind of fungi). Complex life is not a tree of animals, but a tree of protists. Could there really be that many out there? A wonderful YouTube channel, Journey to the Microcosmos, has managed to make these secret stars shine. In beautiful, narrated microscope footage, they reveal an incredible stage most humans never get to see. There is the majestic blue-green Stentor coeruleus, shapeshifting between a trumpet and a sack of marbles, regenerating after an injury. Lacrymaria olor, literally “tears of a swan,” stretches its incredibly long and whip-like neck to hunt prey. An unidentified ciliate slowly loses its shape and life, as its cell membrane — a thin border between order and chaos — gives way.
There’s more to this than eye candy. Once considered genetic bycatch on the journey to multicellularity, Protista actually dominate the medley of complex life.
III. Elusive Folds
The extreme difficulty of categorizing protists has caused our understanding of them to lag. Last November, DeepMind — the same group that previously trained artificial intelligence programs to beat humans in Go and StarCraft — successfully applied their expertise to predicting how proteins fold inside cells, a problem which has stumped scientists for decades. However, even their algorithm struggles to fold protist proteins.
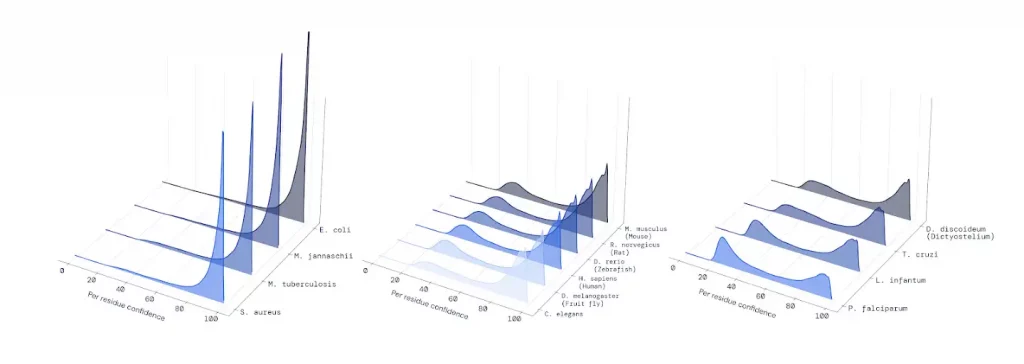
There are obvious reasons for this. The first step of the AlphaFold algorithm is to search DNA databases for similar versions of a protein in other organisms, because those relatives can contain information about its structure. Amino acids that appear to mutate at the same time, for example, might be close together inside a protein. The ability to predict how a protein folds can depend on how many related sequences can be found.
Hundreds of thousands of bacterial and archaeal genomes are publicly available, and AlphaFold’s confidence reflects this wealth of information. Meanwhile, there have been around a thousand sequencing projects for protists, the majority of which are parasites, and their proteins remain an enigma. We will know more soon: last year marked the birth of the Protist 10,000 Genomes Project, which joins similar sequencing campaigns for plants, fungi, arthropods, dogs, birds, and fish, and prokaryotes in the microbiome.
The Gordon and Betty Moore Foundation have also invested in high-risk projects for genetic tool development in marine protists. To accelerate research, the foundation encouraged the exchange of protocols and failures. At the conclusion of the grants in 2020, research groups described genetic transformation of 13 new species, advancements in 8, and failed attempts in 17.
Protist synthetic biology is finally leaving harbor on its maiden voyage.
IV. Beyond Fermentation
Most of what is successful at scale in synthetic biology today involves coaxing E. coli and yeast to convert sugar into more valuable products. This is usually achieved through fermentation, and we did not arrive here by chance.
A handful of model organisms dominate our awareness of biology. Scientists first studied E. coli because it was fast and easy to grow. We have domesticated baker’s and brewer’s yeast for thousands of years. Fruit flies, peas, nematodes, and mice were not chosen because they were particularly interesting, but because they were easily accessible in the lab environment.
Ideas about what to pursue in synthetic biology are overwhelmingly shaped by a select few organisms held in favor, because attempting to genetically engineer any organism before we understand its biology comprehensively is an agonizing task.
But biological design will not be limited to steel tanks forever. What do we want to do after fermentation? While our understanding of thoroughly-studied model organisms is still far from complete, the knowledge we gained with their help will allow us to choose new non-model organisms with specific intentions. The things we create in the future will be determined by the abilities of the organisms we choose to research today.
Although evolutionary biologists moved beyond Whittaker’s five kingdom system long ago, the idea endures because of its flaws: it is simple, intuitive, and reflects what we think we know. As we have seen, single-celled eukaryotes are not the stepping-off point for animal evolution. They are both highly evolved and compose the majority of complex life, like folds of dark matter among bright stars.
Understanding protists — “animalcules,” as Anton van Leuwenhoek called them — will not only improve our ability to engineer plants, animals, and fungi, but give us a whole new way of looking at life itself.
V. The Last Frontier
Fermentation is preferable to fossil fuel-powered manufacturing, but to reach a net carbon-negative output, the future of biosynthesis will have to be green — literally. Today, plankton produce around half of the oxygen we breathe. Billions of years ago, photosynthesis was stolen from bacteria in a Promethean soup when a eukaryote engulfed a cyanobacterium. Because chloroplasts were once free-living organisms, many retain a functional genome distinct from the cell nucleus.
Chlorella is a spherical alga with the highest density of chlorophyll of any plant. Meanwhile, Chlamydomonas is another promising cell factory — so reliable that there is a genetic engineering kit available for it. Its single, horseshoe-shaped chloroplast is much simpler and more responsive to genetic manipulation than those in plants, making Chlamydomonas a candidate for the radical engineering of photosynthesis. Can we increase its efficiency by replacing key enzymes or enabling nitrogen fixation?
Plants and algae are also machines for storing the energy of sunlight in fuel. Biodiesel today is made from pressing the oil out of soybeans, but if you look at a well-fed diatom under the microscope, it will also contain visible lipid droplets. Although synthetic biology has already gone through its biofuels phase once, they are not obsolete, but still in their infancy.
There are clear implications of efficient photosynthesis in algae, but protists have more to offer. While prokaryotes are genetically diverse — 70% DNA similarity being a good cutoff for bacteria belonging to the same species, the same distance between humans and zebrafish — eukaryotes are structurally diverse. They may share similar underlying metabolisms but have divergent body plans, even at the unicellular level.
Diatoms, radiolarians, and foraminifera are all ancient organisms that secrete highly accurate cell walls. Radiolarians are like the gothic cousins to diatoms, which create symmetric glass boxes — one side slightly smaller and fitting into the other. Imagine micron-level patterned glass for nanobiotechnology. The Great Pyramids of Giza were built from limestone almost entirely saturated by the tiny fossilized shells of foraminifera. When alive, the amoeba-like organism sends a myriad of tentacles through its tough shell to sense the environment around it.
Other candidates for non-model model organisms harbor information that is fundamental or contrary to what we have learned so far. How do Stentor correctly track and repair cell morphology? How does Sterkiella recompile its genome from joining 200,000 tiny lengths of DNA? Naegleria transforms its entire cytoskeleton to transition between crawling and swimming — all without actin, the protein that is usually the primary actor of cell movement.
Finally, we know the least about the viruses that infect protists. There is evidence that a vast array of RNA viruses that infect marine protists dominate the oceans, but only a handful have been characterized. Many tools in our genetic toolbox today, from restriction enzymes to CRISPR, come from the millennia-long arms race between viruses and bacteria. What might have been created in the mysterious viruses and their tiny, eukaryotic hosts during their eternal battle in the oceans?
Consider where microbiology was 25 years ago, after E. coli and yeast were sequenced for the first time, knowing that they are now the bread and butter of synthetic biology. What rules will there be for engineering more complex, eukaryotic life, and what might we have missed?
VI. At once past and future
Witnessing me nerd out above, it probably won’t surprise you to learn that during Covid-19, I decided to get a microscope. I wanted to figure out what was inside my aquarium, find a charming gobe-like algae called Volvox, a cult favorite in introductory biology textbooks, and see the diversity of diatoms.
The experience merits getting your hands dirty. Look down the microscope eyepiece, and there is a moment when things click. You can suspend the disbelief that these molecular machines, with no way of knowing they’re being observed, are really alive. You feel like Anton van Leuwenhoek discovering a rich, three-dimensional, microbial world full of life.
Peering down the microscope is like scrying into the past and the future at the same time.
I have seen predatory ciliates hunting diatoms, held in the weave of some aquatic mycelium, their food vacuoles flashing with iridescence and full of glass. From stagnant, neon-green aquarium water, I have watched banana-shaped algae spin in the tiny currents generated by rotifers. The rotifers flinch and retract when copepods bump into them, then hesitantly elongate to beat their wheel-like circles of cilia, and the algae spin again.
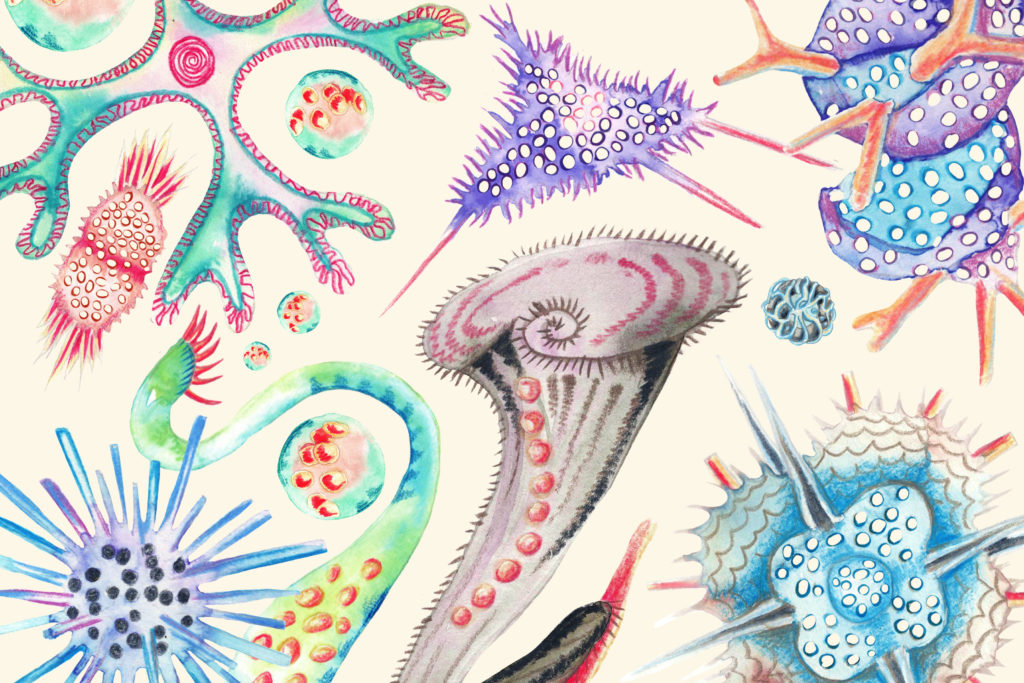
I want there to be more of this, and I want to glimpse what compelled Ernst Haeckel to draw. His most famous work is one portfolio, Kunstformen der Natur, or Art Forms in Nature. The compositions, macabre Radiolaria skeletons and lurid jellyfish tendrils, have inspired art since the twentieth century style of Art Nouveau — defining features of which include curved glass and organic, flowing design.
Peering down the microscope is like scrying into the past and the future at the same time. You see not only the primordial patterns of life, but also what beautiful things we might be able to grow in the future.